A tool for drawing ANI clustermap between all-vs-all microbial genomes
Project description
ANIclustermap
Overview
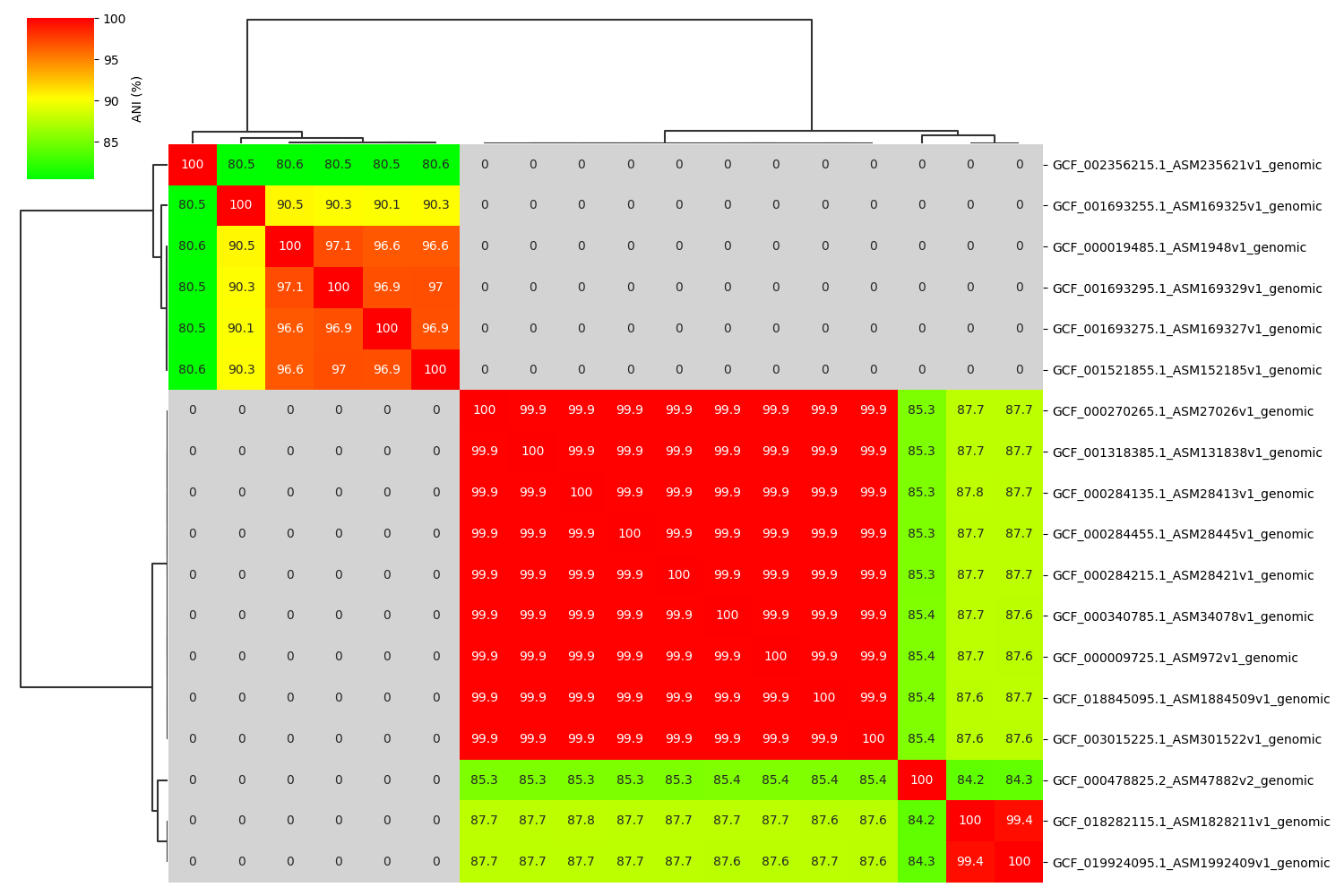
ANIclustermap is easy-to-use tool for drawing ANI(Average Nucleotide Identity) clustermap between all-vs-all microbial genomes.
Fig1. ANIclustermap between all-vs-all 18 genomes. If no similarity detected by fastANI, filled in gray.
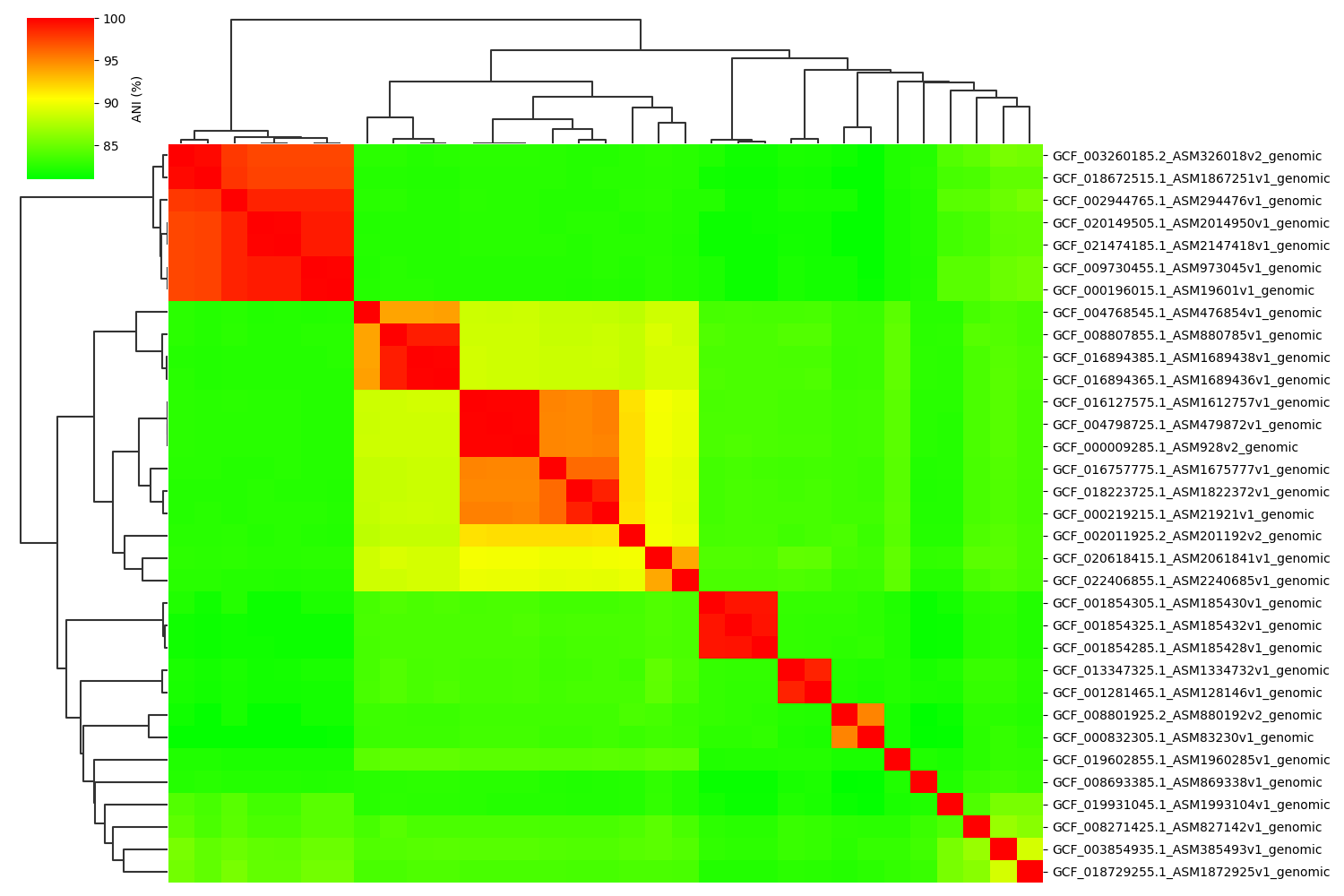
Fig2. ANIclustermap between all-vs-all 33 genomes.
Installation
ANIclustermap is implemented in Python3. fastANI is required to calculate ANI.
Install PyPI stable version with pip:
pip install aniclustermap
Install latest development version with pip:
pip install git+https://github.com/moshi4/ANIclustermap.git
Workflow
Description of ANIclustermap's automated workflow.
- Calculate ANI between all-vs-all microbial genomes by fastANI.
If no similarity detected by fastANI, NA is output. In that case, NA is replaced by 0.0.
If previous result available at the time of re-run, reuse previous result. - Clustering ANI matrix by scipy's UPGMA method.
- Using clustered matrix, draw ANI clustermap by seaborn.
Usage
Basic Command
ANIclustermap -i [Genome fasta directory] -o [output directory]
Options
-h, --help show this help message and exit
-i I, --indir I Input genome fasta directory (*.fa|*.fna[.gz]|*.fasta)
-o O, --outdir O Output directory
-t , --thread_num fastANI thread number parameter (Default: MaxThread - 1)
--fig_width Figure width (Default: 10)
--fig_height Figure height (Default: 10)
--dendrogram_ratio Dendrogram ratio to figsize (Default: 0.15)
--cmap_colors cmap interpolation colors parameter (Default: 'lime,yellow,red')
--cmap_gamma cmap gamma parameter (Default: 1.0)
--annotation Show ANI value annotation (Default: OFF)
-v, --version Print version information
Example Command
ANIclustermap -i ./example/input/small_dataset/ -o ./aniclustermap_result --fig_width 15
Output Contents
ANIclustermap outputs 3 types of files.
-
ANIclustermap.[png|svg](example1, example2)
ANI clustermap result figure. -
ANIclustermap_matrix.tsv(example)
Clustered all-vs-all ANI matrix. -
ANIclustermap_dendrogram.nwk(example)
Newick format clustering dendrogram.
Gallery
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file aniclustermap-1.0.0.tar.gz.
File metadata
- Download URL: aniclustermap-1.0.0.tar.gz
- Upload date:
- Size: 7.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.12 Linux/5.13.0-1017-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d29f02524ad88d39e0d2a63117cc939966de04f25c7f5f1d6e29be28bbcb4ddd
|
|
| MD5 |
75f4bdafe937e4b910c7cc31c5445374
|
|
| BLAKE2b-256 |
c20cc2af5dc4078f1665962cef585d79192dd8e72995be9f755e4f0eedfe51e8
|
File details
Details for the file aniclustermap-1.0.0-py3-none-any.whl.
File metadata
- Download URL: aniclustermap-1.0.0-py3-none-any.whl
- Upload date:
- Size: 288.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.12 Linux/5.13.0-1017-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3bd5fc36ed2f1583b37cab4d995b90a432d28f06205cc1b16c1aef897186e9d1
|
|
| MD5 |
ac4b7e2d88eabc5d854ce8ce891574bf
|
|
| BLAKE2b-256 |
1e664ff1dd82ddeaf5df9c55365a719185d1d4eb0001343ec08e604038875040
|