A Python wrapper for minimap2-rs
Project description
Python bindings for the Rust FFI minimap2 library. In development! Feedback appreciated!
Why?
PyO3 makes it very easy to create Python libraries via Rust. Further, we can use Polars to export results as a dataframe (which can be used as-is, or converted to Pandas). Python allows for faster experimentation with novel algorithms, integration into machine learning pipelines, and provides an opportunity for those not familiar with Rust nor C/C++ to use minimap2.
Current State
Very early alpha. Please use, and open an issue for any features you need that are missing, and for any bugs you find.
How to use
Requirements
Polars and PyArrow, these should be installed when you install minimappers2
Creating an Aligner Instance
aligner = map_ont()
aligner.threads(4)
If you want an alignment performed, rather than just matches, enable .cigar()
aligner = map_hifi()
aligner.cigar()
Please note, at this time the following syntax is NOT supported:
aligner = map_ont().threads(4).cigar()
Creating an index
aligner.index("ref.fa")
To save a built-index, for future processing use:
aligner.index_and_save("ref.fa", "ref.mmi")
Then next time you use the index will be faster if you use the saved index instead.
aligner.load_index("ref.mmi")
Aligning a Single Sequence
query = Sequence(seq_name, seq)
aligner.map1(query)
# Example
seq = "CCAGAACGTACAAGGAAATATCCTCAAATTATCCCAAGAATTGTCCGCAGGAAATGGGGATAATTTCAGAAATGAGAG"
result = aligner.map1(Sequence("MySeq", seq))
Where seq_name and seq are both strings. The output is a Polars DataFrame.
Aligning Multiple Sequences
seqs = [Sequence("name of seq 1", seq1),
Sequence("name of seq 2", seq1)]
result = aligner.map(seqs)
Example Notebook
Please see the example notebook for more examples.
Mapping a file
Please open an issue if you need to map files from this API.
Results
All results are returned as Polars dataframes. You can convert Polars dataframes to Pandas dataframes with .to_pandas()
- Polars is the fastest dataframe library in the Python Ecosystem.
- Polars provides a nice data bridge between Rust and Python.
For more information, please see the Polars User Guide or the Polars Guide for Pandas users.
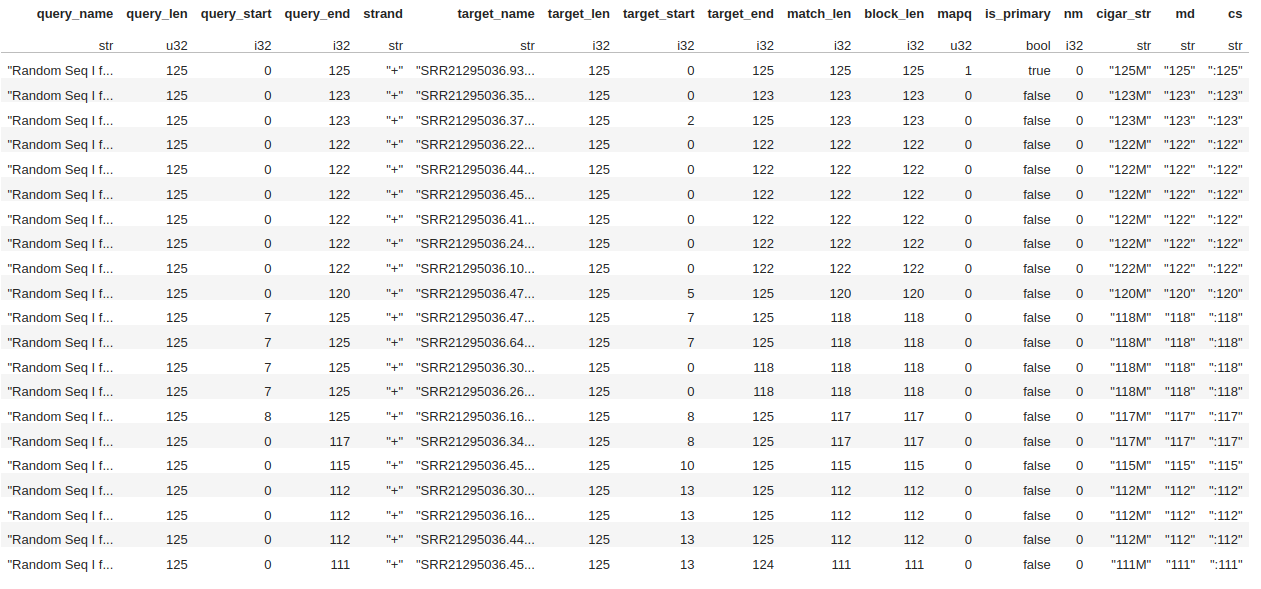
Example of Results
Here is an image of the resulting dataframe
NOTE Mapq, Cigar, and others will not show up unless .cigar() is enabled on the aligner itself.
Errors
As this is a very-early stage library, error checking is not yet implemented. When things crash you will likely need to restart your python interpreter (jupyter kernel). Let me know what happened and open an issue and I will get to it.
Compatability
-
Linux: Yes
-
Mac: Unknown
-
Windows: Unlikely
-
x86_64: Yes
-
aarch64: Unknown (open an issue)
-
neon: No (Open an issue)
-
Google Colab: Yes
Performance
Effort has been made to make this as performant as possible, but if you need more performance, please use minimap2 directly and import the results.
Citation
You should cite the minimap2 papers if you use this in your work.
Li, H. (2018). Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics, 34:3094-3100. [doi:10.1093/bioinformatics/bty191][doi]
and/or:
Li, H. (2021). New strategies to improve minimap2 alignment accuracy. Bioinformatics, 37:4572-4574. [doi:10.1093/bioinformatics/btab705][doi2]
Changelog
0.1.5
- Updated minimap2-rs, polars, pyo3 deps
- Add new presets
0.1.4
- Update pyo3, polars, minimap2-rs, and mimalloc deps
0.1.1
- Update pyo3 and polars deps
- Add with_seq for indexing TODO
0.1.0
- Initial Functions implemented
- Return results as Polars dfs
Funding
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file minimappers2-0.1.7.tar.gz.
File metadata
- Download URL: minimappers2-0.1.7.tar.gz
- Upload date:
- Size: 2.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: maturin/1.8.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c94b6a9b7fa807a3586719ce806a50e1d82800b86f704019f22e8220f12c4e3d
|
|
| MD5 |
f68d253f3ef5ca0c3a169eb8760a3437
|
|
| BLAKE2b-256 |
b22b86702e9300f3883f778941045fcc59ae861f44f3f2ae828a7ff83eba1331
|
File details
Details for the file minimappers2-0.1.7-cp37-abi3-manylinux_2_34_x86_64.whl.
File metadata
- Download URL: minimappers2-0.1.7-cp37-abi3-manylinux_2_34_x86_64.whl
- Upload date:
- Size: 4.2 MB
- Tags: CPython 3.7+, manylinux: glibc 2.34+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: maturin/1.8.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
22af4946fc0b7991a7891daceed8756738e98d3ae9b17ce17ee6b6a09842c708
|
|
| MD5 |
be9a46f98ad21fc3f48602b22f7b0b3d
|
|
| BLAKE2b-256 |
df7e0c98585122bb0c2c844cc2277b3a7fc55b54b70156dc750cafab7290ce08
|