A quality control pipeline for illumina data set. This pipeline removes contaminants (e.g. Phix), performs fastqc, adapter cleaning and trimming and checks for contaminants
Project description
This is is the quality_control pipeline from the Sequana project
- Overview:
A quality control pipeline for illumina data set. This pipeline removes contaminants (e.g. Phix), performs fastqc, adapter cleaning and trimming and checks for contaminants
- Input:
Raw fastq files
- Output:
Cleaned fastQ files, remove phix and adapters + taxonomy
- Status:
production.
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
Installation
You must install Sequana first:
pip install sequana
Then, just install this package:
pip install sequana_quality_control
Usage
sequana_pipelines_quality_control --help sequana_pipelines_quality_control --input-directory DATAPATH
This creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd quality_control sh quality_control.sh # for a local run
This launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the pipeline itself and its configuration files and then execute the pipeline yourself with specific parameters:
snakemake -s quality_control.rules -c config.yaml --cores 4 --stats stats.txt
Or use sequanix interface.
Requirements
This pipelines requires the following executable(s):
fastqc
bwa
kraken (optional)
krona (optional)
sambamba
samtools
pigz
cutadapt [or atropos]
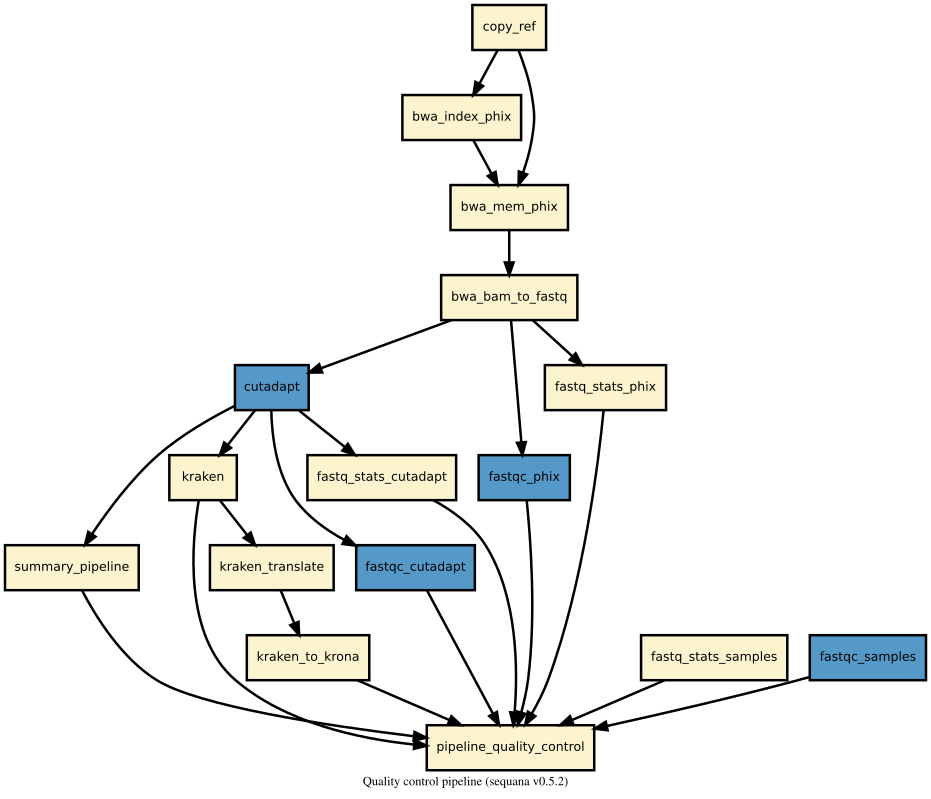
Details
This pipeline runs quality_control in parallel on the input fastq files (paired or not). A brief sequana summary report is also produced.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
ChangeLog
Version |
Description |
|---|---|
0.8.2: |
|
0.8.1: |
uses more sequana tools to handle the options |
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file sequana_quality_control-0.8.2.tar.gz.
File metadata
- Download URL: sequana_quality_control-0.8.2.tar.gz
- Upload date:
- Size: 490.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.15.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/40.4.3 requests-toolbelt/0.9.1 tqdm/4.39.0 CPython/3.5.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2cfe7c77d4749d3dd90c3749aefb71e96bf60252ead3dc4dccd814dd1862ead8
|
|
| MD5 |
7bb35c8773d767e41a1baaacb8dd2ee5
|
|
| BLAKE2b-256 |
73888fd427a50a2ce11b202e3b8d712ba5c8a45c4a2c6e877a6c08e5f4676f61
|











