Fragment Quality Control for ATAC-seq
Project description
ATACFragQC
The Python toolkits designed to control the fragment quality of Bulk/SingCell ATAC-seq.
Installation
python3 -m pip install ATACFragQC
Usage
# Basic usage
ATACFragQC [options] -i <input.bam> -r <reference.gtf>
# For more information
ATACFragQC -h
Features
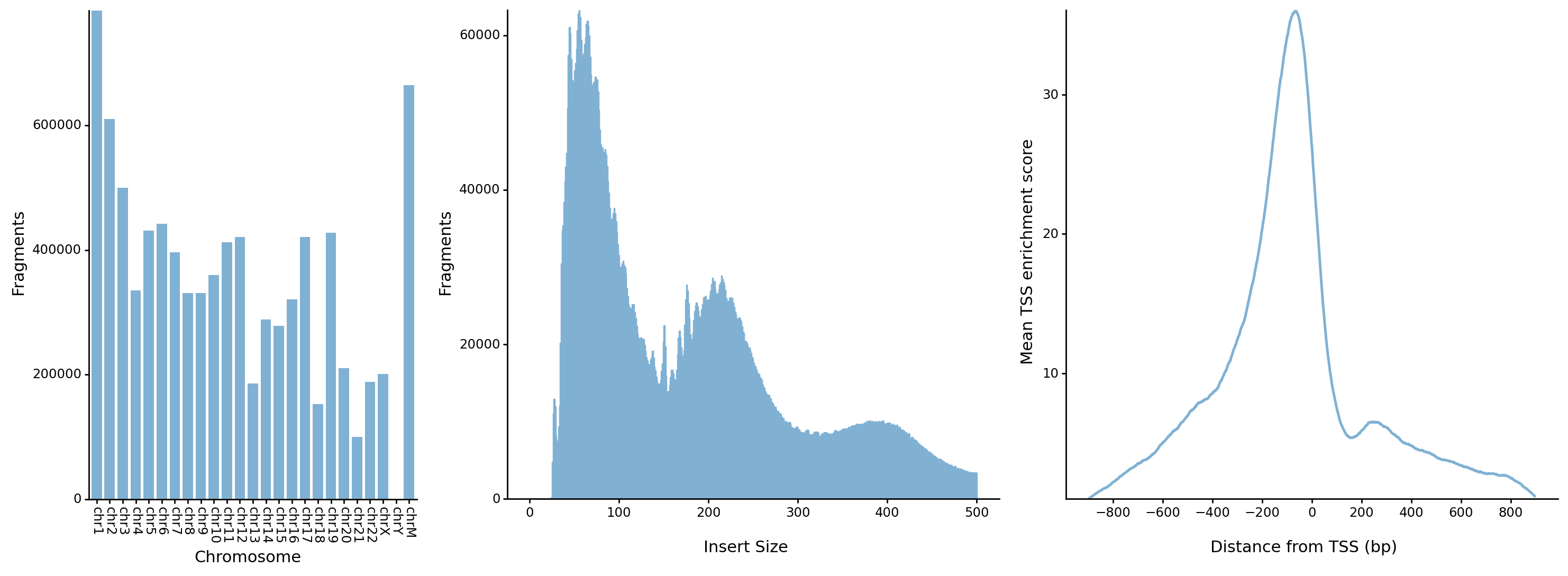
- The distrubution of fragments in chromosomes
- The distrubution of fragment lengths
- The distrubution of fragments around transcription start sites (TSSs)
- Other feature would be supported in the future ...
Overview
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
ATACFragQC-0.2.3.tar.gz
(5.0 kB
view details)
File details
Details for the file ATACFragQC-0.2.3.tar.gz.
File metadata
- Download URL: ATACFragQC-0.2.3.tar.gz
- Upload date:
- Size: 5.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.23.0 setuptools/49.3.1 requests-toolbelt/0.9.1 tqdm/4.45.0 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8ad2f8a3c9c47ab17c4dcd4c4449cda935de0943c41fae35eb3fbea2ddebaec4
|
|
| MD5 |
8102376dc45fcf80c8ecda8992369c19
|
|
| BLAKE2b-256 |
e96f336e4b84189ef5b84ce0fa434d516698a9a74c15c0dbc94f30d7b8f0babc
|